Hyperbolic mixed, dual modification (activation → inhibition)
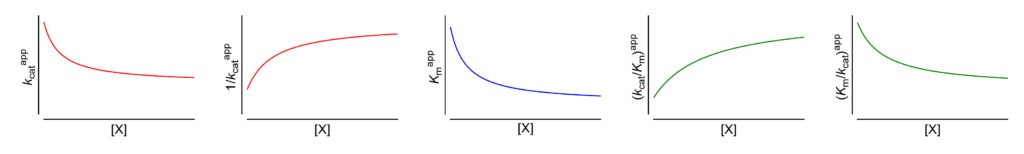
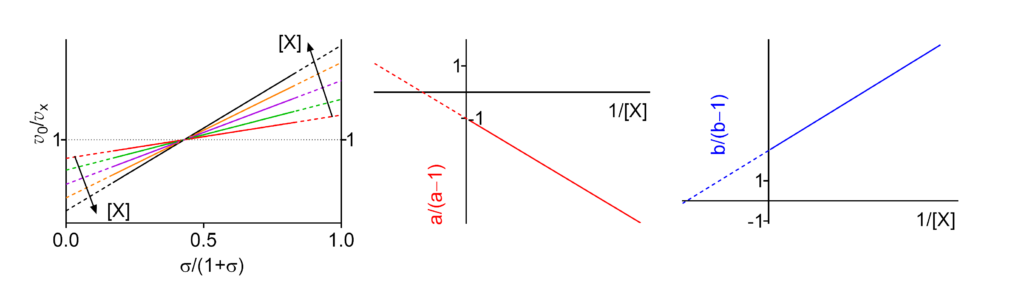
Featured examples
| # | Enzyme Species | EC no. | Modifier | Substrate(1) | Name given by authors(2) | Reference (3) |
|---|---|---|---|---|---|---|
| 1 | Cathepsin K Homo sapiens | 3.4.22.38 | Chondroitin sulfate | Cbz-Phe-Arg-7-amino-4-methylcoumarylamide | Hyperbolic mixed inhibition + nonessential activation α = 0.13, β = 0.85, KX = 33 μM, σ 0 = 4.8 | Novinec (2010) |
| 2 | Cathepsin K Homo sapiens | 3.4.22.38 | Dextran sulfate | Cbz-Phe-Arg-7-amino-4-methylcoumarylamide | Hyperbolic mixed inhibition + nonessential activation α = 0.29, β = 0.50, KX = 18 μM, σ 0 = 0.45 | Novinec (2010) |
| 3 | Cathepsin K Homo sapiens | 3.4.22.38 | Heparin | Cbz-Phe-Arg-7-amino-4-methylcoumarylamide | Hyperbolic mixed inhibition + nonessential activation (4) α1 = 0.16, β1 = 0.93, KX,1 = 0.78 μM α2 = 0.29, β2 = 0.50, KX,2 = 3.8 μM | Novinec (2010) |
| 4 | Coagulation factor IXa Homo sapiens | 3.4.21.22 | ISIS 2302 (5) | CH3-SO2-(D)-CHG-Gly-Arg-pNA | Partial, uncompetitive inhibition (6) α = 0.17, β = 0.26, KX = 87.7 nM, σ 0 = 0.12 | Sheehan (2001) |
| 5 | 3-Phosphoshikimate 1-carboxyvinyltransferase Bacillus subtilis | 2.5.1.19 | Glyphosate | Shikimate-3-phosphate (with 100 mM NH4Cl) | Mixed-type inhibition (7) | Fischer (1987) |
| 6 | D-amino acid oxidase Homo sapiens | 1.4.3.3 | Risperidone | D-Alanine | Activation/inhibition (8) | Abou El-Magd (2010) |
| 7 | Arachidonate 15-lipoxygenase Homo sapiens | 1.13.11.33 | 13-(S)-HODE | Arachidonic acid | K-type activation + V-type inhibition α = 0.2, β = 0.76, KX = 1.2 μM, σ 0 = 2.3 | Joshi (2013) |
| 8 | Laccase Trametes versicolor | 1.10.3.2 | Arsenate (As5+) | 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid) | Mixed inhibition with preference toward an uncompetitive inhibition Km↓, V↓, KX = 15.4 μM | Wang (2016) |
| 9 | Lipopolysaccharide heptosyltransferase I Escherichia coli | 2.4.99.B6 | Hexene-β-D-glucopyranoside | ADP-L-glycero-D-manno-heptose | Noncompetitive (mixed-competitive) inhibition (9) | Nkosana (2018) |
(1) Always the varied substrate. In two- or more-substrate reactions the concentration(s) of the non varied substrate(s) is/are kept constant.
(2) Name of the mechanism given by the authors in the quoted reference. α, β and the inhibition/activation constants for the modifier (X), uniformly denoted KX, are the values specified by the authors. In some cases, missing parameters have been calculated from graphical or tabular data provided in the papers. In two- or more-substrate reactions, KX represents an apparent constant at given concentrations of the fixed substrates and no calculations of the intrinsic values have been attempted.
(3) Full references at the end of the page provide also the digital object identifier (doi), if available. Clicking the authors (highlighted) opens the reference in PubMed.
(4) Complex kinetic mechanisms that cannot be detailed here, please consult the original publication.
(5) The 20-mer phosphorothioate oligodeoxyribonucleotide 5′-GCCCAAGCTGGCATCCGTCA-3′.
(6) Only inhibition was observed because the range of substrate concentrations was larger than the critical substrate concentration of 0.17 mM.
(7) Commenting that activation occurs at low [S] and inhibition at higher [S].
(8) The authors commented: ‘There may be some doubt whether to call ‘I’ an activator or an inhibitor as it clearly activates at low substrate concentrations, but inhibits at higher substrate concentrations’.
(9) Too few data (each three of [S] and [X]) to calculate KX, α and β with sufficient precision. However, the dependencies of the kinetic parameters on [X] were investigated from the data in Table 2 of the paper and were sufficient to diagnose the mechanism.
References
- Abou El-Magd RM, Park HK, Kawazoe T, Iwana S, Ono K, Chung SP, Miyano M, Yorita K, Sakai T, Fukui K (2010) The effect of risperidone on D-amino acid oxidase activity as a hypothesis for a novel mechanism of action in the treatment of schizophrenia. J Psychopharmacol 24: 1055-1067. doi:10.1177/0269881109102644
- Fischer RS, Rubin JL, Gaines CG, Jensen RA (1987) Glyphosate sensitivity of 5-enol-pyruvylshikimate-3-phosphate synthase from Bacillus subtilis depends upon state of activation induced by monovalent cations. Arch Biochem Biophys 256: 325-334. doi:10.1016/0003-9861(87)90453-X
- Joshi N, Hoobler EK, Perry S, Diaz G, Fox B, Holman TR (2013) Kinetic and structural investigations into the allosteric and pH effect on the substrate specificity of human epithelial 15-lipoxygenase-2. Biochemistry 52: 8026-8035. doi:10.1021/bi4010649
- Nkosana NK, Czyzyk DJ, Siegel ZS, Cote JM, Taylor EA (2018) Synthesis, kinetics and inhibition of Escherichia coli Heptosyltransferase I by monosaccharide analogues of Lipid A. Bioorg Med Chem Lett 28: 594-600. doi:10.1016/j.bmcl.2018.01.040
- Novinec M, Kovačič L, Lenarčič B, Baici A (2010) Conformational flexibility and allosteric regulation of cathepsin K. Biochem J 429: 379-389. doi:10.1042/BJ20100337
- Sheehan JP, Phan TM (2001) Phosphorothioate oligonucleotides inhibit the intrinsic tenase complex by an allosteric mechanism. Biochemistry 40: 4980-4989. doi:10.1021/bi002396x
- Wang T, Milton RD, Abdellaoui S, Hickey DP, Minteer SD (2016) Laccase inhibition by arsenite/arsenate: Determination of inhibition mechanism and preliminary application to a self-powered biosensor. Anal Chem 88: 3243-3248. doi:10.1021/acs.analchem.5b04651