Hyperbolic mixed, predominantly specific inhibition
Featured examples
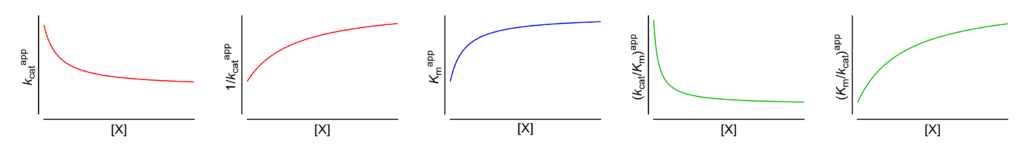
- Km↑ The apparent Michaelis constant increases with increasing [X]
- V ↓ (∴kcat ↓) The apparent limiting rate, and therefore the catalytic constant, decrease with increasing [X]
- (V/Km)↓ (∴kcat/Km ↓) The apparent V/Km ratio, and therefore the specificity constant, decrease with increasing [X]
These symbols are shown only when the featured dependencies of the parameters on modifier concentration have been demonstrated by the authors.
| # | Enzyme Species | EC no. | Modifier | Substrate(1) | Name given by authors (2) | Reference(3) |
|---|---|---|---|---|---|---|
| 1 | Stratum corneum chymotryptic enzyme Homo sapiens | 3.4.21.117 | Antileukoproteinase (4) | MeO-Succinyl-Arg-Pro-Tyr- p-nitroanilide | Hyperbolic mixed-type inhibition α = 5.6, β = 0.16, KX = 63 nM | Franzke (1996) |
| 2 | Non-specific serine/threonine protein kinase Rattus norvegicus | 2.7.11.1 | Heparin fragment (6-mer) | Peptide RRRADDSDDDDD | Hyperbolic partial non-competitive mixed type inhibition Parameters inconsistent (5) | O'Farrel (1999) |
| 3 | Leukocyte elastase Homo sapiens | 3.4.21.37 | Gold thiomalate | t-Butyloxycarbonyl-alanyl-p-nitrophenyl ester | Hyperbolic mixed-type inhibition α = 1.6, β = 0.6, KX = 33 μM | Baici (1981) |
| 4 | Coagulation factor Xa Homo sapiens | 3.4.21.6 | Antistasin (6) | CH3SO2-Leu-Gly-Arg- p-nitroanilide | Mixed, primarily competitive inhibition KX= 0.3-0.6 nM (6) α ≈ 1.3, β ≈ 0.4, KX ≈ 0.3 nM | Dunwiddie (1989) |
| 5 | Leukocyte elastase Homo sapiens | 3.4.21.37 | Suramin | Suc-Ala-Ala-Ala-p-nitroanilide | Partial inhibition α = 4.0, β = 0.43, KX = 0.18 μM | Cadène (1997) |
| 6 | Cathepsin G Homo sapiens | 3.4.21.20 | Suramin | Suc-Ala-Ala-Pro-Phe- p-nitroanilide | Partial inhibition α = 2.3, β = 0.29, KX = 0.08 μM | Cadène (1997) |
| 7 | Myeloblastin Homo sapiens | 3.4.21.76 | Suramin | MeO-Suc-Ala-Ala-Pro-Val-thiobenzylester | Partial inhibition α = 1.8, β = 0.63, KX = 0.5 μM | Cadène (1997) |
| 8 | Leukocyte elastase Homo sapiens | 3.4.21.37 | Dermatan sulfate Mr= 31,000; 62 DU/chain | MeO-Suc-Ala-Ala-Pro-Val-p-nitroanilide | Hyperbolic mixed, predominantly competitive inhibition (7) α = 4.2, β = 0.23 K0.5= 0.25 μMDU, h = 1.1 | Kostoulas (1997) |
| 9 | Leukocyte elastase Homo sapiens | 3.4.21.37 | Chondroitin 4-sulfate Mr =18,000; 36 DU/chain | MeO-Suc-Ala-Ala-Pro-Val-p-nitroanilide | Hyperbolic mixed, predominantly competitive inhibition (7) α = 3.1, β = 0.09 K0.5= 45 μMDU, h = 1.1 | Kostoulas (1997) |
| 10 | Leukocyte elastase Homo sapiens | 3.4.21.37 | Chondroitin 6-sulfate Mr = 29,000; 58 DU/chain | MeO-Suc-Ala-Ala-Pro-Val-p-nitroanilide | Hyperbolic mixed, predominantly competitive inhibition (7) α = 2.1, β = 0.59 K0.5= 0.16 μMDU, h = 1.6 | Kostoulas (1997) |
| 11 | Leukocyte elastase Homo sapiens | 3.4.21.37 | Dermatan sulfate low-Mr Mr = 10,200; 20 DU/chain | MeO-Suc-Ala-Ala-Pro-Val-p-nitroanilide | Hyperbolic mixed, predominantly competitive inhibition (7) α = 2.2, β = 0.85 K0.5= 1.1 μMDU, h = 1.8 | Kostoulas (1997) |
| 12 | Leukocyte elastase Homo sapiens | 3.4.21.37 | Chondroitin 4,6-disulfate Mr = 27,600; 46 DU/chain | MeO-Suc-Ala-Ala-Pro-Val-p-nitroanilide | Hyperbolic mixed, predominantly competitive inhibition (7) α = 2.4, β = 0.34 K0.5= 0.07 μMDU, h = 2.9 | Kostoulas (1997) |
| 13 | Leukocyte elastase Homo sapiens | 3.4.21.37 | Dermatan 4,6-disulfate Mr = 25,000; 41 DU/chain | MeO-Suc-Ala-Ala-Pro-Val-p-nitroanilide | Hyperbolic mixed, predominantly competitive inhibition (7) α = 2.0, β = 0.24 K0.5= 0.07 μMDU, h = 2.5 | Kostoulas (1997) |
| 14 | Leukocyte elastase Homo sapiens | 3.4.21.37 | Chondroitin N,4-disulfate Mr = 6,900; 12 DU/chain | MeO-Suc-Ala-Ala-Pro-Val-p-nitroanilide | Hyperbolic mixed, predominantly competitive inhibition (7) α = 2.4, β = 0.52 K0.5= 0.4 μMDU, h = 1.7 | Kostoulas (1997) |
| 15 | Leukocyte elastase Homo sapiens | 3.4.21.37 | Dermatan N,4-disulfate Mr = 6,600; 12 DU/chain | MeO-Suc-Ala-Ala-Pro-Val-p-nitroanilide | Hyperbolic mixed, predominantly competitive inhibition (7) α = 3.2, β = 0.53 K0.5= 0.7 μMDU, h = 1.9 | Kostoulas (1997) |
| 16 | Caspase-2 Homo sapiens | 3.4.22.55 | AR_F8 (8) | Acetyl-Val-Asp-Val-Ala-Asp-7-amino-4-methylcoumarylamide | Hyperbolic mixed, predominantly competitive inhibition α = 2.9, β = 0.04, KX = 0.29 nM | Schweizer (2007) |
| 17 | ADAMTS 13 endopeptidase (9) Homo sapiens | 3.4.24.87 | cVWF63 (10) | Synthetic peptide (11) | Hyperbolic mixed type inhibition α = 5.4, β = 0.12, KX = 0.44 μM | Gao (2006) |
| 18 | Acetylcholinesterase Electrophorus electricus | 3.1.1.7 | Meldola blue | Acetylthiocholine | Hyperbolic mixed inhibition α = 1.9, β = 0.28, KX = 0.035 μM | Küçükkilinç (2007) |
| 19 | Acetylcholinesterase Electrophorus electricus | 3.1.1.7 | Nile blue | Acetylthiocholine | Hyperbolic mixed inhibition α = 1.8, β = 0.15, KX = 0.026 μM | Küçükkilinç (2007) |
| 20 | Acetylcholinesterase Electrophorus electricus | 3.1.1.7 | Methylene blue | Acetylthiocholine | Hyperbolic mixed inhibition α = 2.4, β = 0.18, KX = 0.017 μM | Küçükkilinç (2007) |
| 21 | Acetylcholinesterase Electrophorus electricus | 3.1.1.7 | Gallamine triethiodide (without acetonitrile) | Acetylthiocholine | Mixed-type inhibition with pronounced competitive component α = 15, β = 0.25, KX = 270 μM | Pietsch (2009) |
| 22 | Acetylcholinesterase Electrophorus electricus | 3.1.1.7 | Compound 11 | Acetylthiocholine | Hyperbolic mixed-type inhibition α = 2.0, β = 0.14, KX = 1.3 μM | González-Tanarro (2011) |
| 23 | Acetylcholinesterase Electrophorus electricus | 3.1.1.7 | Compound 12 | Acetylthiocholine | Hyperbolic mixed-type inhibition α = 1.5, β = 0.17, KX = 1.9 μM | González-Tanarro (2011) |
| 24 | Acetylcholinesterase Electrophorus electricus | 3.1.1.7 | Compound 13 | Acetylthiocholine | Hyperbolic mixed-type inhibition α = 1.6, β = 0.16, KX = 2.4 μM | González-Tanarro (2011) |
| 25 | Acetylcholinesterase Electrophorus electricus | 3.1.1.7 | Compound 14 | Acetylthiocholine | Hyperbolic mixed-type inhibition α = 1.6, β = 0.12, KX = 1.5 μM | González-Tanarro (2011) |
| 26 | Aminopeptidase N Homo sapiens | 3.4.11.2 | Cholic acid | Leu-Gly | Hyperbolic mixed-type inhibition α = 2.5, β = 0.33, KX = 0.91 mM | Nakanishi (1989) |
| 27 | Aminopeptidase N Homo sapiens | 3.4.11.2 | Chenodeoxycholic acid | Leu-Gly | Hyperbolic mixed-type inhibition α = 2.7, β = 0.13, KX = 0.24 mM | Nakanishi (1989) |
| 28 | Spermidine synthase Bos taurus | 2.5.1.16 | Spermidine | Putrescine | Hyperbolic mixed-type inhibition (12) Kic = 10-110 μM, Kiu > 500 μM | Raina (1984) |
| 29 | Spermidine synthase Bos taurus | 2.5.1.16 | Spermidine | Decarboxylated adenosylmethionine | Hyperbolic mixed-type inhibition (12) Kic = 80-160 μM, Kiu > 500 μM | Raina (1984) |
| 30 | Peroxiredoxin Homo sapiens | 1.11.1.15 | 4-tert-butylcatechol | H2O2 | Partial mixed noncompetitive inhibition α = 2.0, β = 0.77, KX = 0.09 mM | Chow (2016) |
| 31 | Alpha,alpha trehalase Chironomus riparius | 3.2.1.28 | 6-epi-casuarine | Trehalose | Hyperbolic mixed-type inhibition α = 4.8, β = 0.4, KX = 48 nM | D'Adamio (2015) |
| 32 | Alpha,alpha trehalase Chironomus riparius | 3.2.1.28 | 7-deoxycasuarine | Trehalose | Hyperbolic mixed-type inhibition α = 2.5, β = 0.5, KX = 810 nM | D'Adamio (2015) |
| 33 | Ethanolamine-phosphate phospho-lyase Homo sapiens | 4.2.3.2 | Phosphate (pH = 8.8) | O-phosphoethanolamine | Hyperbolic mixed-type inhibition α = 8.5, β = 0.48, KX = 8 mM | Schiroli (2015) |
| 34 | Cathepsin K Homo sapiens | 3.4.22.38 | Compound 1 | Cbz-Phe-Arg-7-amino-4-methylcoumarylamide | Hyperbolic mixed, predominantly competitive inhibition α = 6.7, β = 0.55, KX = 190 μM | Novinec (2014) |
| 35 | Cathepsin K Homo sapiens | 3.4.22.38 | Compound 2 | Cbz-Phe-Arg-7-amino-4-methylcoumarylamide | Hyperbolic mixed, predominantly competitive inhibition α = 7.2, β = 0.74, KX = 150 μM | Novinec (2014) |
| 36 | Cathepsin K Homo sapiens | 3.4.22.38 | Compound 3 | Cbz-Phe-Arg-7-amino-4-methylcoumarylamide | Hyperbolic mixed, predominantly competitive inhibition α = 1.7, β = 0.44, KX = 110 μM | Novinec (2014) |
| 37 | Sterol esterase Sus scrofa domestica | 3.1.1.13 | Compound 4a | p-Nitrophenyl butyrate | Hyperbolic mixed-type inhibition α = 5.5, β = 0.13, KX = 1.22 μM | Heng (2011) |
| 38 | Cholinesterase Homo sapiens | 3.1.1.8 | Serotonin | o-Nitrophenylacetate | Hyperbolic competitive inhibition α = 1.9, β = 0.6, KX = 1.7 mM | Masson (2008) |
| 39 | Acetylcholinesterase Electrophorus electricus | 3.1.1.7 | Methyl green | Acetylthiocholine | Hyperbolic mixed inhibition α = 2.7, β = 0.20, KX = 0.27 μM | Küçükkilinç (2008) |
| 40 | Linoleate 13S-lipoxygenase Glycine max | 1.13.11.12 | Oleyl sulfate | Linoleic acid | Hyperbolic mixed-type inhibition α = 4.6, β = 0.85, KX = 0.7 μM Km↑, kcat↓, kcat/Km↓ | Mogul (2000) |
| 41 | Cytochrome P450 (13) Rattus norvegicus | —(14) | Primaquine | Aminopyrine | Slope-hyperbolic, intercept-hyperbolic noncompetitive inhibition α = 2.05, β = 0.54, KX = 21 μM | Murray (1986) |
| 42 | Coagulation factor XIa Homo sapiens | 3.4.21.27 | Coagulation factor IX | L-Pyroglutamyl-L-prolyl-L-arginine-p-nitroanilide | Hyperbolic mixed inhibition α = 2.7, β = 0.50, KX = 0.22 μM | Ogawa (2005) |
| 43 | Coagulation factor XIa Homo sapiens | 3.4.21.27 | Active site-inhibited factor IXa (factor IXai) | L-Pyroglutamyl-L-prolyl-L-arginine-p-nitroanilide | Hyperbolic mixed inhibition α = 2.5, β = 0.9, KX = 0.11 μM | Ogawa (2005) |
| 44 | Cholinesterase Homo sapiens (native enzyme) | 3.1.1.8 | Tosyllysine chloromethyl ketone | Butyrylthiocholine | Hyperbolic mixed inhibition α = 41.6, β = 0.065, KX = 39 μM | Cengiz (1997) |
| 45 | Cholinesterase Homo sapiens (native enzyme) | 3.1.1.8 | Tosylphenylalanine chloromethyl ketone | Butyrylthiocholine | Hyperbolic mixed inhibition α = 10.8, β = 0.26, KX = 16 μM | Cengiz (1997) |
| 46 | Cholinesterase Homo sapiens (heat desensitized enzyme, 24 h, 45°C) | 3.1.1.8 | Tosyllysine chloromethyl ketone | Butyrylthiocholine | Hyperbolic mixed inhibition α = 3.9, β = 0.52, KX = 17 μM | Çokuğraş (2004) |
| 47 | Coagulation factor VIIa Homo sapiens | 3.4.21.21 | Peptide A-183 EEWEVLCWTWETCER | Coagulation factor X | Partial mixed-type inhibition α = 2.6, β = 0.27, KX ≈ 200 pM | Dennis (2001) |
| 48 | Cellulose 1,4-beta-cellobiosidase Trichoderma reesei | 3.2.1.91 | Cellobiose (disaccharide, reaction product) | Cellulose | Mixed hyperbolic inhibition (14) α = 37.9, β = 0.055, KX = 29 μM (cellobiose present initially) | Bezerra (2011) |
| 49 | Cellulose 1,4-beta-cellobiosidase Trichoderma reesei | 3.2.1.91 | Cellobiose (disaccharide, reaction product) | Cellulose | Mixed hyperbolic inhibition (14) α = 51.7, β = 0.063, KX = 29 μM (no cellobiose present initially) | Bezerra (2011) |
| 50 | Papain Carica papaya | 3.4.22.1 | Fukugetin | Cbz-Phe-Arg-7-amino-4-methylcoumarylamide | Hyperbolic mixed inhibition α = 1.8, β = 0.12, KX = 13.5 μM | Assis (2013) |
| 51 | Acetolactate synthase Escherichia coli | 2.2.1.6 | Valine | Pyruvate | Mixed V/K type inhibition α = 3.2, β = 0.62, KX = 4.6 μM | Mendel (2001) |
| 52 | ABC-type quaternary amine transporter(15) Homo sapiens | 7.6.2.9 | Tiagabine | γ-Aminobutyric acid | Mixed competitive/non-competitive inhibition α ≈ 8, β ≈ 0.2, KX ≈ 2 μM (16) | Kragholm (2013) |
| 53 | Serine racemase Homo sapiens | 5.1.1.18 | 1,4-dihydronicotinamide mononucleotide | L-Serine | Hyperbolic mixed-type inhibition α = 5, β = 0.8, KX = 18 μM | Bruno (2016) |
| 54 | Cathepsin B Homo sapiens | 3.4.22.1 | DARPin 8h6 | Cbz-Arg-Arg-7-amino-4-methylcoumarylamide | Mixed-mode inhibition with a very strong competitive component α = 6.6, β = 0.15, KX = 34 pM (tight-binding) | Kramer (2017) |
| 55 | Cathepsin B Mus musculus | 3.4.22.1 | DARPin 8h6 | Cbz-Arg-Arg-7-amino-4-methylcoumarylamide | Mixed-mode inhibition with a very strong competitive component α = 6.1, β = 0.06, KX = 35 pM (tight-binding) | Kramer (2017) |
| 56 | Leukocyte elastase Homo sapiens | 3.4.21.37 | Ethylenediamine-tetraacetic acid (EDTA) | MeO-Suc-Ala-Ala-Pro-Val-p-nitroanilide | Hyperbolic mixed inhibition with partial competitive and partial noncompetitive components (17) α = 2.1, β = 0.26, KX = 6.6 mM | Tyagi (1991) |
| 57 | Receptor protein-tyrosine kinase Homo sapiens | 2.7.10.1 | Lavendustin A | ATP | Hyperbolic mixed-type inhibition (18) α > 1, β < 1, KX ≤ 1 nM | Hsu (1991) |
| 58 | Phosphoserine phosphatase Mycobacterium tuberculosis | 3.1.3.3 | L-serine | O-phospho-L-serine | Hyperbolic mixed, predominantly specific inhibition (19) α = 3.5, β = 0.0006, KX = 22 μM | Pierson (2023) |
(1) Always the varied substrate. In two- or more-substrate reactions the concentration(s) of the non varied substrate(s) is/are kept constant.
(2) Name of the mechanism given by the authors in the quoted reference. α, β and the inhibition/activation constants for the modifier (X), uniformly denoted KX, are the values specified by the authors. In some cases, missing parameters have been calculated from graphical or tabular data provided in the papers. In two- or more-substrate reactions, KX represents an apparent constant at given concentrations of the fixed substrates and no calculations of the intrinsic values have been attempted.
(3) Full references at the end of the page provide also the digital object identifier (doi), if available. Clicking the authors (highlighted) opens the reference in PubMed.
(4) Antileukoproteinase, or secretory leukocyte proteinase inhibitor, is a human endogenous inhibitor of serine peptidases.
(5) The results in Fig. 4 clearly demonstrate the HMx(Sp>Ca)I mechanism, whereas α = 0.88 and β = 0.013 (Table 2) are characteristic of HMx(Sp<Ca)I; Ki = 19 μM. The discrepancy is possibly due to the fitting procedure with intertwined parameters.
(6) Antistasin, a protein from the salivary glands of the Mexican leech Haementeria officinalis. It selectively inhibits Factor Xa acting as anticoagulant. This is an uncommon case of slow-onset inhibition described in ‘Kinetics of enzyme-modifier interactions‘ by A. Baici, pp. 408-414. The approximate values of α, β and KX were calculated from Fig. 3 (left) in the paper.
(7) One or more enzyme molecules can bind to a single chain of the glycosaminoglycans in a cooperative manner. The equation used to calculate the inhibition parameters was: vi = v0 -{(v0 – v∞)[X]h}/{K0.5h + [X]h}, where vi, v0 and v∞ are the initial velocities in the absence of modifier, in its presence and at saturating modifier concentration, respectively. X is the modifier, K0.5 is the inhibitor concentration for which the velocity equals (v0 – v∞)/2 and h is the Hill coefficient that is usually not an integer. All glycosaminoglycans (the inhibitors) were oligodisperse with known molecular mass, composition and number of disaccharide units (DU) on the chain. Their concentration and K0.5 are expressed as micromoles of DU × L-1 (μMDU). The electrostatically-driven interaction between elastase and glycosaminoglycans depends on ionic composition and ionic strength. The data in this table refer to a Tris/HCl buffer, pH = 7.40, ionic strength = 100 mM and 25°C.
(8) AR_F8 = designed ankyrin repeat protein specifically targeted to human caspase-2. Tight-binding with human caspase-2.
(9) Recombinant ADAMTS 13 truncated after the spacer domain.
(10) cVWF63 = C-terminal cleavage product of Von Willebrand factor.
(11) Synthetic peptide in which leucine at position 3 is isotopically doubly-labeled with (13)C and (15)N (GP([(13)C(15)N]L)GSDREQAPNLVY.
(12) Product inhibition. Hyperbolic inhibition clearly demonstrated (α > 1, β < 1 ) but the value of β was not shown.
(13) An ‘aminopyrine N-demethylase activity’ in a rat hepatic microsomal fraction, possibly due to ‘CYP2C19, EC 1.14.14.1 (unspecific monooxygenase) or another cytochrome P450 activity.
(14) Assays conducted with complete progress curves to which the integrated Michaelis -Menten equation was fitted. Models comprising substrate inhibition and parabolic inhibition by the concomitant binding of two inhibitor molecules to the enzyme were considered. However, by applying the Akaike information criterion, a simpler model equivalent to the general modifier mechanism was selected as the most probable.
(15) Transporters belong to the EC-enzyme class 7, translocases, added in 2018 to the other 6 enzyme classes. This paper by Kragholm et al. contains a kinetic study of a human ABC-type quaternary amine transporter commonly known as betaine/GABA transporter 1, a target for the treatment of epilepsy. Besides the inhibitor featured here, also N-(1-benzyl-4-piperidinyl)-2,4-dichlorobenzamide was investigated as inhibitor, see under LMx(Sp<Ca)I.
(16) Approximate calculations from the data in the paper.
(17) The same mechanism is shared with the chelators QUIN-2 and FURA-2.
(18) Notwithstanding a difficult method for measuring the kinetic courses of the reactions, the authors did a great job in providing sound experimental evidence to assert that the mechanism is of the tight-binding, slow-onset inhibition type. Moreover, the data in Fig. 6 are consistent with hyperbolic mixed, predominantly specific inhibition as the basic mechanism. The kinetic parameters are approximate, but epitomize very well the goal of this elegant study.
(19) The authors conducted an impeccable kinetic analysis in identifying the basic modifier mechanism in a complex system. HMx(Sp>Ca)I, an allosteric modifier, can be confused with the corresponding linear counterpart LMx(Sp>Ca)I because, as can be seen in some examples in the table above (see the studies 9, 44, 49, and 55) the parameter β is often very small. Only an accurate examination of the saturation curves in the presence of sufficiently high concentrations of the inhibitor can reveal the hyperbolic nature of this inhibition mechanism.
References
- Assis DM, Gontijo VS, de Oliveira Pereira I, Santos JA, Camps I, Nagem TJ, Ellena J, Izidoro MA, dos Santos Tersariol IL, de Barros NM, Doriguetto AC, dos Santos MH et al. (2013) Inhibition of cysteine proteases by a natural biflavone: behavioral evaluation of fukugetin as papain and cruzain inhibitor. J Enzyme Inhib Med Chem 28: 661-670. doi:10.3109/14756366.2012.668539
- Baici A, Salgam P, Fehr K, Böni A (1981) Inhibition of human elastase from polymorphonuclear leucocytes by gold sodium thiomalate and pentosan polysulfate (SP-54). Biochem Pharmacol 30: 703-708. doi:10.1016/0006-2952(81)90154-4
- Bezerra RMF, Dias AA, Fraga I, Pereira AN (2011) Cellulose hydrolysis by cellobiohydrolase Cel7A shows mixed hyperbolic product inhibition. Appl Biochem Biotechnol 165: 178-189. doi:10.1007/s12010-011-9242-y
- Bruno S, Marchesani F, Dellafiora L, Margiotta M, Faggiano S, Campanini B, Mozzarelli A (2016) Human serine racemase is allosterically modulated by NADH and reduced nicotinamide derivatives. Biochem J 473: 3505-3516. doi:10.1042/BCJ20160566
- Cadène M, Duranton J, North A, Sitahar M, Chignard M, Bieth JG (1997) Inhibition of neutrophil serine proteinases by suramin. J Biol Chem 272: 9950-9955. doi:10.1074/jbc.272.15.9950
- Cengiz D, Çokuğraş AN, Kilinç K, Tezcan EF (1997) Histidine modification of human serum butyrylcholinesterase. Biochem Mol Med 61: 52-57. doi:10.1006/bmme.1997.2578
- Chow ML, Troussicot L, Martin M, Doumèche B, Guillière F, Lancelin J-M (2016) Predicting and understanding the enzymatic inhibition of human peroxiredoxin 5 by 4-substituted pyrocatechols by combining funnel metadynamics, solution NMR, and steady-state kinetics. Biochemistry 55: 3469-3480. doi:10.1021/acs.biochem.6b00367
- Çokuğraş AN, Cengiz D, Tezcan EF (2004) Do alkylating agents modify the histidine residue of the desensitized butyrylcholinesterase? Protein J 23: 495-500. doi:10.1007/s10930-004-7876-0
- D’Adamio G, Sgambato A, Forcella M, Caccia S, Parmeggiani C, Casartelli M, Parenti P, Bini D, Cipolla L, Fusi P, Cardona F (2015) New synthesis and biological evaluation of uniflorine A derivatives: towards specific insect trehalase inhibitors. Org Biomol Chem 13: 886-892. doi:10.1039/C4OB02016B
- Dennis MS, Roberge M, Quan C, Lazarus RA (2001) Selection and characterization of a new class of peptide exosite inhibitors of coagulation factor VIIa. Biochemistry 40: 9513-9521. doi:10.1021/bi010591l
- Dunwiddie C, Thornberry NA, Bull HG, Sardana M, Friedman PA, Jacobs JW, Simpson E (1989) Antistasin, a leech-derived inhibitor of factor Xa. Kinetic analysis of enzyme inhibition and identification of the reactive site. J Biol Chem 264: 16694-16699.
- Franzke CW, Baici A, Bartels J, Christophers E, Wiedow O (1996) Antileukoprotease inhibits stratum corneum chymotryptic enzyme. Evidence for a regulative function in desquamation. J Biol Chem 271: 21886-21890. doi:10.1074/jbc.271.36.21886
- Gao WQ, Anderson PJ, Majerus EM, Tuley EA, Sadler JE (2006) Exosite interactions contribute to tension-induced cleavage of von Willebrand factor by the antithrombotic ADAMTS13 metalloprotease. Proc Natl Acad Sci USA 103: 19099-19104. doi:10.1073/pnas.0607264104
- González-Tanarro CM, Gütschow M (2011) Hyperbolic mixed-type inhibition of acetylcholinesterase by tetracyclic thienopyrimidines. J Enzyme Inhib Med Chem 26: 350-358. doi:10.3109/14756366.2010.504674
- Heng S, Tieu W, Hautmann S, Kuan K, Pedersen DS, Pietsch M, Gütschow M, Abell AD (2011) New cholesterol esterase inhibitors based on rhodanine and thiazolidinedione scaffolds. Bioorg Med Chem 19: 7453-7463. doi:10.1016/j.bmc.2011.10.042
- Hsu CY, Persons PE, Spada AP, Bednar RA, Levitzki A, Zilberstein A (1991) Kinetic analysis of the inhibition of the epidermal growth factor receptor tyrosine kinase by lavendustin-A and its analogue. J Biol Chem 266: 21105-21112
- Kostoulas G, Hörler D, Naggi A, Casu B, Baici A (1997) Electrostatic interactions between human leukocyte elastase and sulfated glycosaminoglycans: physiological implications. Biol Chem 378: 1481-1489. doi:10.1515/bchm.1997.378.12.1481
- Kragholm B, Kvist T, Madsen KK, Jørgensen L, Vogensen SB, Schousboe A, Clausen RP, Jensen AA, Bräuner-Osborne H (2013) Discovery of a subtype selective inhibitor of the human betaine/GABA transporter 1 (BGT-1) with a non-competitive pharmacological profile. Biochem Pharmacol 86: 521-528. doi:10.1016/j.bcp.2013.06.007
- Kramer L, Renko M, Završnik J, Turk D, Seeger MA, Vasiljeva O, Grütter MG, Turk V, Turk B (2017) Non-invasive in vivo imaging of tumour-associated cathepsin B by a highly selective inhibitory DARPin. Theranostics 7: 2806-2821. doi:10.7150/thno.19081
- Küçükkilinç T, Özer I (2007) Multi-site inhibition of human plasma cholinesterase by cationic phenoxazine and phenothiazine dyes. Arch Biochem Biophys 461: 294-298. doi:10.1016/j.abb.2007.02.029
- Küçükkilinç TT, Ozer I (2008) Inhibition of electric eel acetylcholinesterase by triarylmethane dyes. Chem-Biol Interact 175: 309-311. doi:10.1016/j.cbi.2008.05.008
- Masson P, Froment MT, Gillon E, Nachon F, Lockridge O, Schopfer LM (2008) Kinetic analysis of effector modulation of butyrylcholinesterase-catalysed hydrolysis of acetanilides and homologous esters. FEBS J 275: 2617-2631. doi:10.1111/j.1742-4658.2008.06409.x
- Mendel S, Elkayam T, Sella C, Vinogradov V, Vyazmensky M, Chipman DM, Barak Ze (2001) Acetohydroxyacid synthase: A proposed structure for regulatory subunits supported by evidence from mutagenesis1. J Mol Biol 307: 465-477. doi:10.1006/jmbi.2000.4413
- Mogul R, Johansen E, Holman TR (2000) Oleyl sulfate reveals allosteric inhibition of soybean lipoxygenase-1 and human 15-lipoxygenase. Biochemistry 39: 4801-4807. doi:10.1021/bi992805t
- Murray M, Farrell GC (1986) Mechanistic aspects of the inhibition of microsomal drug oxidation by primaquine. Biochem Pharmacol 35: 2149-2155. doi:10.1016/0006-2952(86)90585-X
- Nakanishi M, Moriyama A, Narita Y, Sasaki M (1989) Aminopeptidase-M from human liver. II. Kinetic analysis of inhibition of the enzyme by bile acids. J Biochem 106: 826-830. doi:10.1093/oxfordjournals.jbchem.a122938
- Novinec M, Lenarčič B, Baici A (2014) Probing the activity modification space of the cysteine peptidase cathepsin K with novel allosteric modifiers. PLoS One 9: e106642. doi:10.1371/journal.pone.0106642
- O’Farrell F, Loog M, Janson IM, Ek P (1999) Kinetic study of the inhibition of CK2 by heparin fragments of different length. Biochim Biophys Acta 1433: 68-75. doi:10.1016/S0167-4838(99)00147-8
- Ogawa T, Verhamme IM, Sun MF, Bock PE, Gailani D (2005) Exosite-mediated substrate recognition of factor IX by factor XIa. The factor XIa heavy chain is required for initial recognition of factor IX. J Biol Chem 280: 23523-23530. doi:10.1074/jbc.M500894200
- Pierson E, De Pol F, Fillet M, Wouters J (2023) A morpheein equilibrium regulates catalysis in phosphoserine phosphatase SerB2 from Mycobacterium tuberculosis. Commun Biol 6: 1024. doi:10.1038/s42003-023-05402-z
- Pietsch M, Christian L, Inhester T, Petzold S, Gütschow M (2009) Kinetics of inhibition of acetylcholinesterase in the presence of acetonitrile. FEBS J 276: 2292-2307. doi:10.1111/j.1742-4658.2009.06957.x
- Raina A, Hyvönen T, Eloranta T, Voutilainen M, Samejima K, Yamanoha B (1984) Polyamine synthesis in mammalian tissues. Isolation and characterization of spermidine synthase from bovine brain. Biochem J 219: 991-1000. doi:10.1042/bj2190991
- Schiroli D, Ronda L, Peracchi A (2015) Kinetic characterization of the human O-phosphoethanolamine phospho-lyase reveals unconventional features of this specialized pyridoxal phosphate-dependent lyase. FEBS J 282: 183-199. doi:10.1111/febs.13122
- Schweizer A, Roschitzki-Voser H, Amstutz P, Briand C, Gulotti-Georgieva M, Prenosil E, Binz HK, Capitani G, Baici A, Plückthun A, Grütter MG (2007) Inhibition of caspase-2 by a designed ankyrin repeat protein: Specificity, structure, and inhibition mechanism. Structure 15: 625-636. doi:10.1016/j.str.2007.03.014
- Tyagi SC, Simon SR (1991) Interaction of neutrophil elastase with hydrophobic polyanionic chelators. Biochem Cell Biol 69: 624-629. doi:10.1139/o91-092